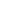Brett J. Baker
Assistant Professor of Marine Science, University of Texas at Austin
Microorganisms are key mediators in nearly all of the planet’s elemental cycles. However, our understanding of the ecological roles of many groups of microbes has been hampered by low-resolution analytical approaches to studying the staggering diversity present in nature. As a result the tree of life is full of branches, which remain undiscovered, and those, which have only been identified in single-gene sequencing surveys (Baker and Dick, 2013). This is a fundamental gap in our understanding of biology. Filling in the genomic gaps in the tree of life will provide a rich context to understand the evolution of life on the planet and will provide us with a genetic understanding of how microbial communities drive biogeochemical cycles.
Recent advances in DNA sequencing technologies and computational analyses have made it possible to reconstruct the genomes and transcriptomes of uncultured natural populations (Baker et al. 2010, 2012 and 2013). I have been involved in the development (Dick et al. 2009) and implementation of environmental omics since the beginning. I was involved in the first metaproteomic study of a microbial community (Ram et al. 2005) and have been using these approaches to track fine-scale evolutionary processes (Denef et al. 2010). Using these techniques I discovered deeply branching, novel groups of microbes (Archaea referred to as ARMAN) that are close to the predicted lower size limit of an organism (Baker et al. 2006). Obtaining complete genomes of the ARMAN phylum revealed that they have signatures of inter-species interactions and form connections to other species in nature (Baker et al. 2010).
More recently, my laboratory has reconstructed the genomes of hundreds of widespread, uncultured sediment microbes to understand how ecological roles are partitioned in these microbial communities. Many of the genomes belong to phyla which have no previous genomic representation and discovered three new groups of bacteria they play important roles in the global carbon cycle (Baker et al. 2015; Lazar, et al, Environ Micro). One of the new branches for which we have obtained several genomes for is a deeply branched member (Thorarchaeota) (Seitz et al. 2016). These genomes have provided rich insights into the evolutionary histories of life on the planet and we have been able to map the flow of carbon and energy, a microbial food web, through sediments with unprecedented detail (Baker et al. 2015).
- Market Data