The US military has released previously classified photos and films related to unidentified flying object (UFO) sightings, which mostly show something blurry moving strangely. Still, I hear that a friend of a friend has gone from thinking there’s a 1% chance that UFOs are aliens to now believing it is 50%. Is he rational?
People are constantly seeing things in the sky they don’t understand. The vast majority are aeroplanes, satellites, weather balloons, clouds, rocket launches, auroras, optical reflections and so on. But for some sightings, there’s no known explanation. The problem is that people jump to the conclusion “unknown = aliens”. And when you think about it, this is fairly odd. Why not angels?
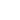
Anyway, I like to do maths instead. The Bayes formula (below), a mainstay of statistics, gives the probability (Pr) of something, given some evidence.

Author provided
Spelled out, it says that the probability that UFOs are aliens given some evidence is equal to how likely it is that the evidence would appear if UFOs really were aliens, times how likely it is that there are aliens. That needs to be divided by how likely the actual evidence is, which is notoriously difficult to work out.
But what we are really interested in is if the evidence tells us we should believe in aliens compared to not believing in aliens. We can do this by dividing the equation above with the counterpart for UFOs not being aliens:

Author provided
When we do this, we also get rid of that pesky factor for how probable the evidence is. The equation now shows how likely it is that UFOs are aliens compared to how likely it is that they are not – after looking at the footage. The result will be one if the options are equally likely, and high if aliens are the stronger bet. It tells us how we should update our beliefs based on new evidence.
There are two factors in the equation. One (second bracket) is how likely we think aliens are. Some might say 50:50, making this factor one, while others may make it very low, like 10-23. This is a statement of belief based on knowledge of the world (using for example the famous Drake equation).
This needs to be multiplied by another factor (first bracket), often called the Bayes factor. It denotes how specific the evidence we see is for aliens v no aliens. If I meet a little green blob claiming to be from Epsilon Eridani, that is relatively specific (but could still somewhat be explained by a prank or me being mad). In this case, the factor may be much bigger than 1 and I get to shift towards thinking there are aliens.
If I see a mysterious blob of light in the sky that could be aliens but could also be a lot of other things, then the factor would not be much different from 1 – the evidence is as specific for aliens as it is for no aliens, and I don’t get much change in belief.
In other words, specificity is hugely important. Weird and unknown things may happen, but if the lights could equally well be faeries, intrusions from the fifth dimension, swamp gas, Chinese drones, sapient octopuses, or anything else, the Bayes factor will still be close to 1. That the world is strange is not evidence for aliens.
My verdict
The latest UFO revelations from the US government doesn’t make me update in the direction of aliens much. Sure, there is lots of weird footage. But it could be explained by many other things: there are no green blobs demanding to be taken to our leader. There’s not even a photo of an alien. Given that earlier research also has made me think the universe is pretty empty, I end up with a very low personal probability estimate of UFOs being aliens.
Here’s my calculation. I start with assuming that aliens visiting is pretty unlikely – I place it somewhere around one in a billion. Why? Because I think the probability of intelligent life per planet is really low, and if there were any out there, it would probably spread on a cosmic scale. Indeed, that we haven’t been paved over already is an important piece of evidence.
As for the specificity of the evidence, I accept that weird things show up, but none of it looks particular for aliens. So my Bayes factor is at best 2 or so (and I think that is too much, actually). So I end up giving a one in 500 million chance to UFOs being aliens after looking at the footage.
One should, however, recognise the great uncertainty here: that one in a billion estimate is based on arguments that could be wrong and are debatable.
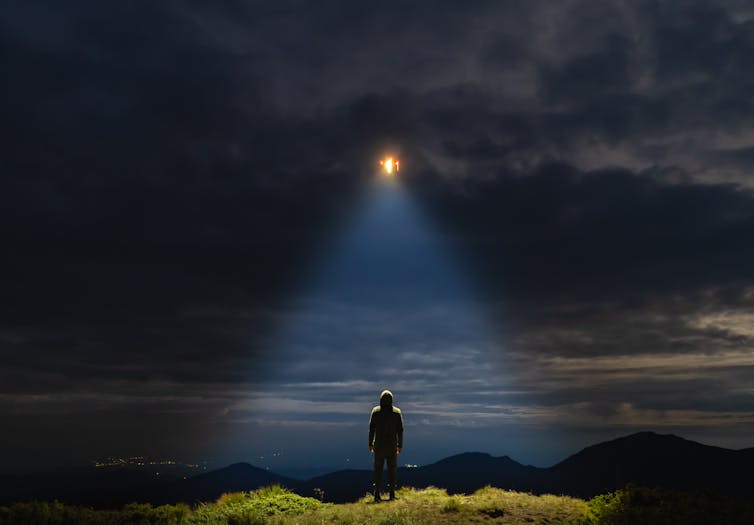

Specific enough? WeAre/Shutterstock
Now imagine I see every TV channel showing footage of a green blob demanding an audience with the UN Secretary General. If it was a real alien, the probability of the footage would be 1. But the probability that it is a super-elaborate prank or that I had a psychotic break is maybe 1 in 1,000 (psychosis is far more common than many think). So by dividing 1 by 1/000, I would get a Bayes factor of 1,000 – boosting my estimate by a factor of 1,000. When I then multiply that, per the equation, by the 1 in a billion probability of aliens visiting, I get a total probability of two in a million.
This would not be enough to think it must be real. But it would be alarming enough to check if my friends are seeing the same thing. Surely they can’t all go mad at the same time – that would be even less likely. If they agree I would boost my estimate by a few more orders of magnitude, to maybe 1/10. I would also check for evidence that it isn’t a super-prank.
As for the current evidence, what would convince me otherwise? More specific evidence, not just blurry lights moving apparently fast. Science did not believe in meteorites until trustworthy, multiple witnesses brought in rocks found to be unknown minerals (a good Bayes factor), and our understanding of the solar system allowed for asteroids.
I suspect actual evidence for visits from extraterrestrial intelligence will be hard to miss. Trying to explain away the weakness of current evidence as aliens being cleverly stealthy does not make them more likely since it makes the evidence unspecific. The search will no doubt go on, but we should look for specific things, not blurry ones.


 Trump Administration to Launch Autism Initiatives Targeting Acetaminophen Use and New Treatment Options
Trump Administration to Launch Autism Initiatives Targeting Acetaminophen Use and New Treatment Options  NASA Partners with Katalyst to Save Swift Observatory with Innovative Docking Mission
NASA Partners with Katalyst to Save Swift Observatory with Innovative Docking Mission  FDA Pilot Program Eases Rules for Nicotine Pouch Makers
FDA Pilot Program Eases Rules for Nicotine Pouch Makers  Neuren Pharmaceuticals Surges on U.S. Patent Win for Rare Disorder Drug
Neuren Pharmaceuticals Surges on U.S. Patent Win for Rare Disorder Drug  Lost in space: MethaneSat failed just as NZ was to take over mission control – here’s what we need to know now
Lost in space: MethaneSat failed just as NZ was to take over mission control – here’s what we need to know now  Neuralink Plans High-Volume Brain Implant Production and Fully Automated Surgery by 2026
Neuralink Plans High-Volume Brain Implant Production and Fully Automated Surgery by 2026  SpaceX Prioritizes Moon Mission Before Mars as Starship Development Accelerates
SpaceX Prioritizes Moon Mission Before Mars as Starship Development Accelerates  SpaceX Pivots Toward Moon City as Musk Reframes Long-Term Space Vision
SpaceX Pivots Toward Moon City as Musk Reframes Long-Term Space Vision  Blue Origin’s New Glenn Achieves Breakthrough Success With First NASA Mission
Blue Origin’s New Glenn Achieves Breakthrough Success With First NASA Mission